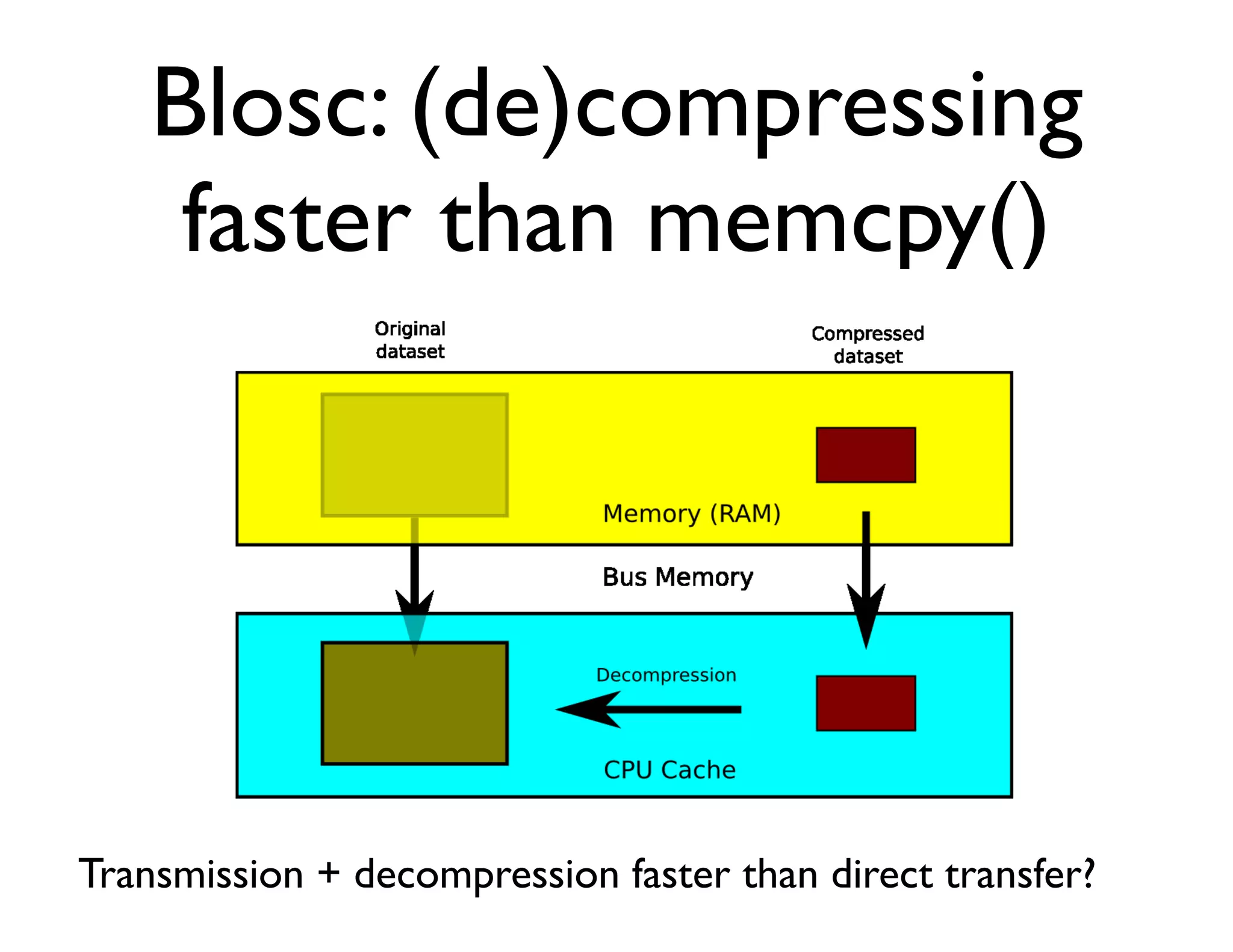
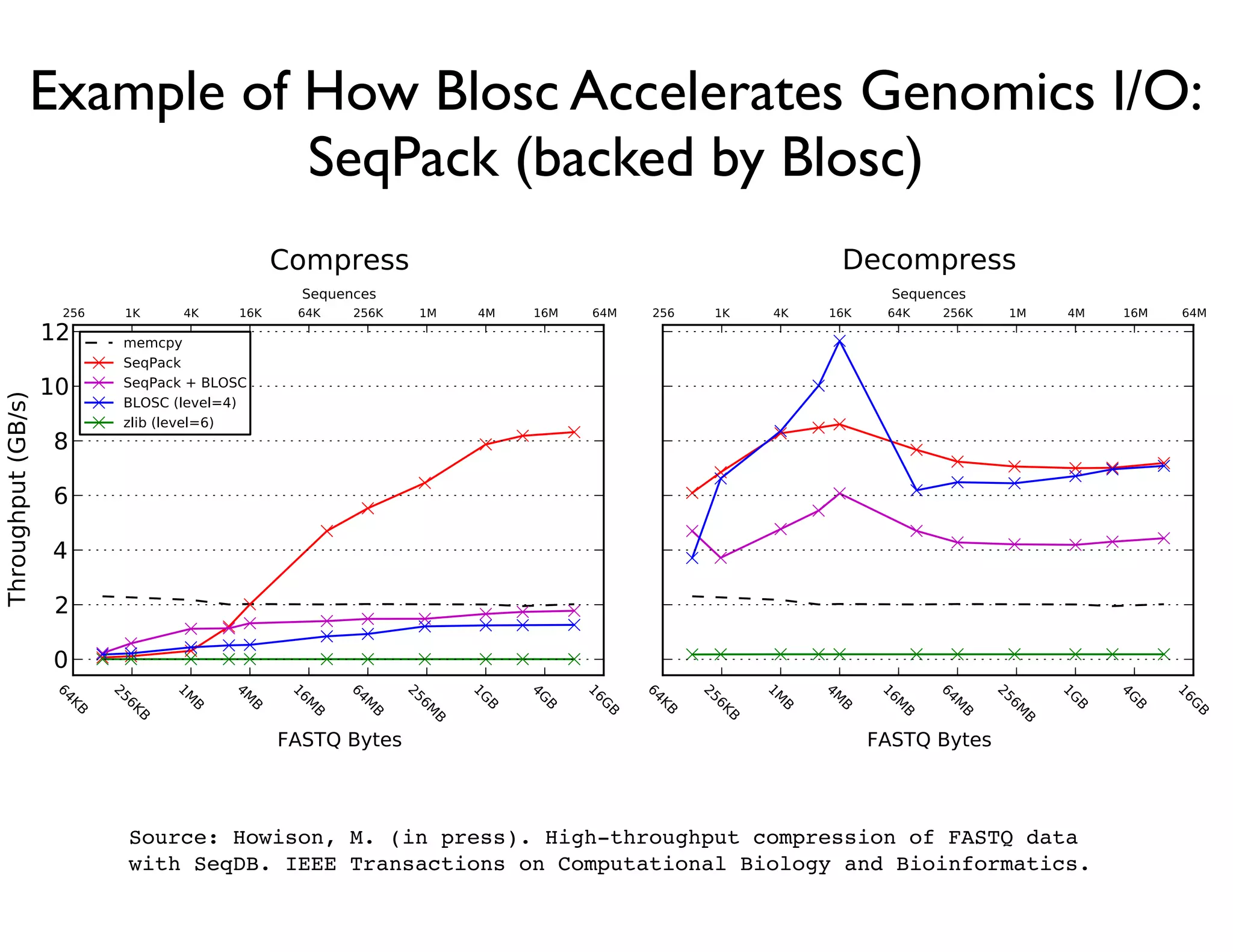
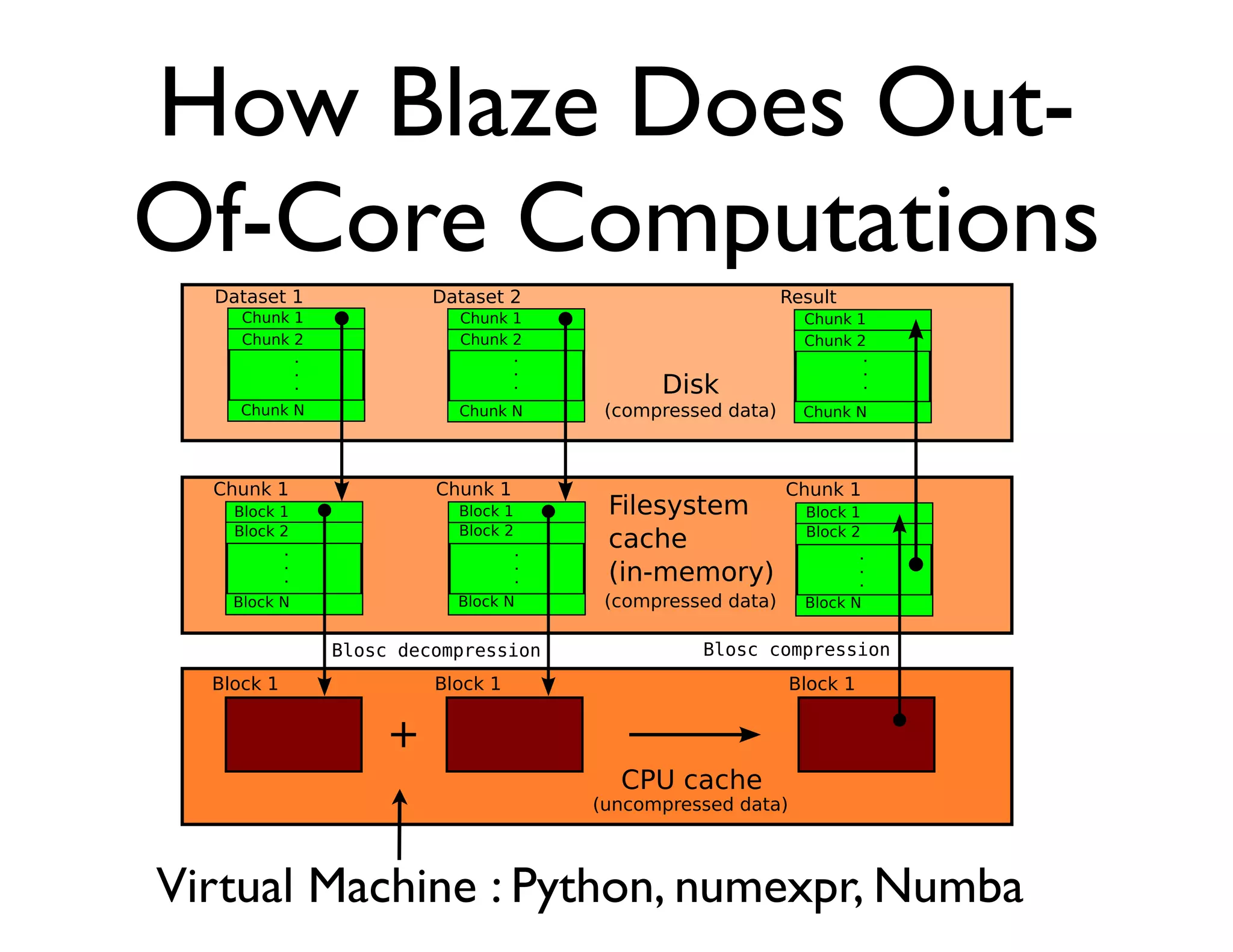
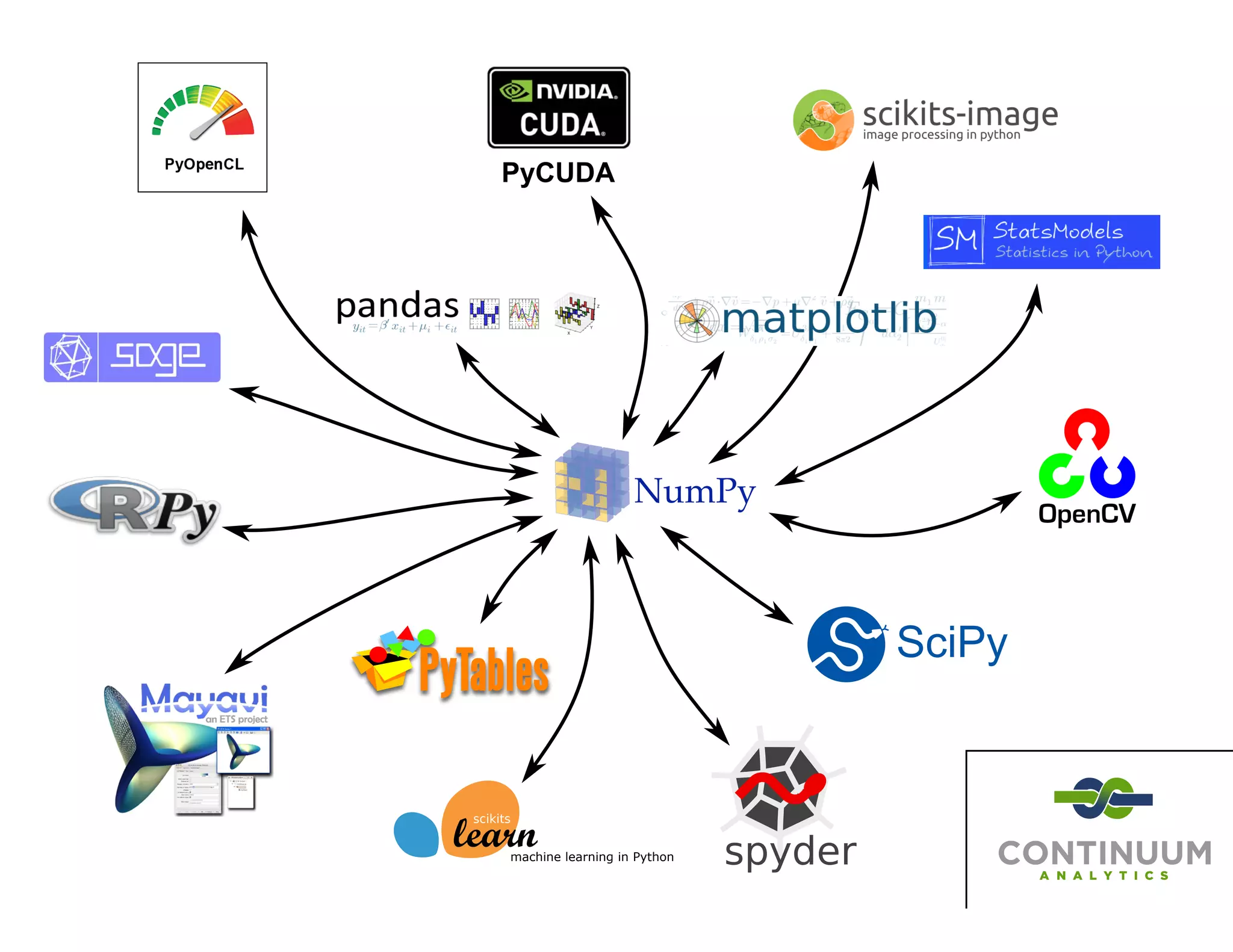
The document discusses the significance of memory access in handling big data, emphasizing that optimal data structures and containers are crucial for performance. It highlights the advantages of using Python and libraries like NumPy and Numba for data computation, illustrating the efficiency improvements through examples and comparisons. The presentation concludes by stressing the importance of selecting appropriate data containers for effective big data management.








![Operating
with NumPy
• array[2]; array[1,1:5, :]; array[[3,6,10]]
• (array1**3 / array2) - sin(array3)
• numpy.dot(array1, array2): access to
optimized BLAS (*GEMM) functions
• and much more...](https://image.slidesharecdn.com/francescalted-howilearnedtostopworryingaboutcpuspeed-121128024051-phpapp01/75/Memory-efficient-applications-FRANCESC-ALTED-at-Big-Data-Spain-2012-10-2048.jpg)


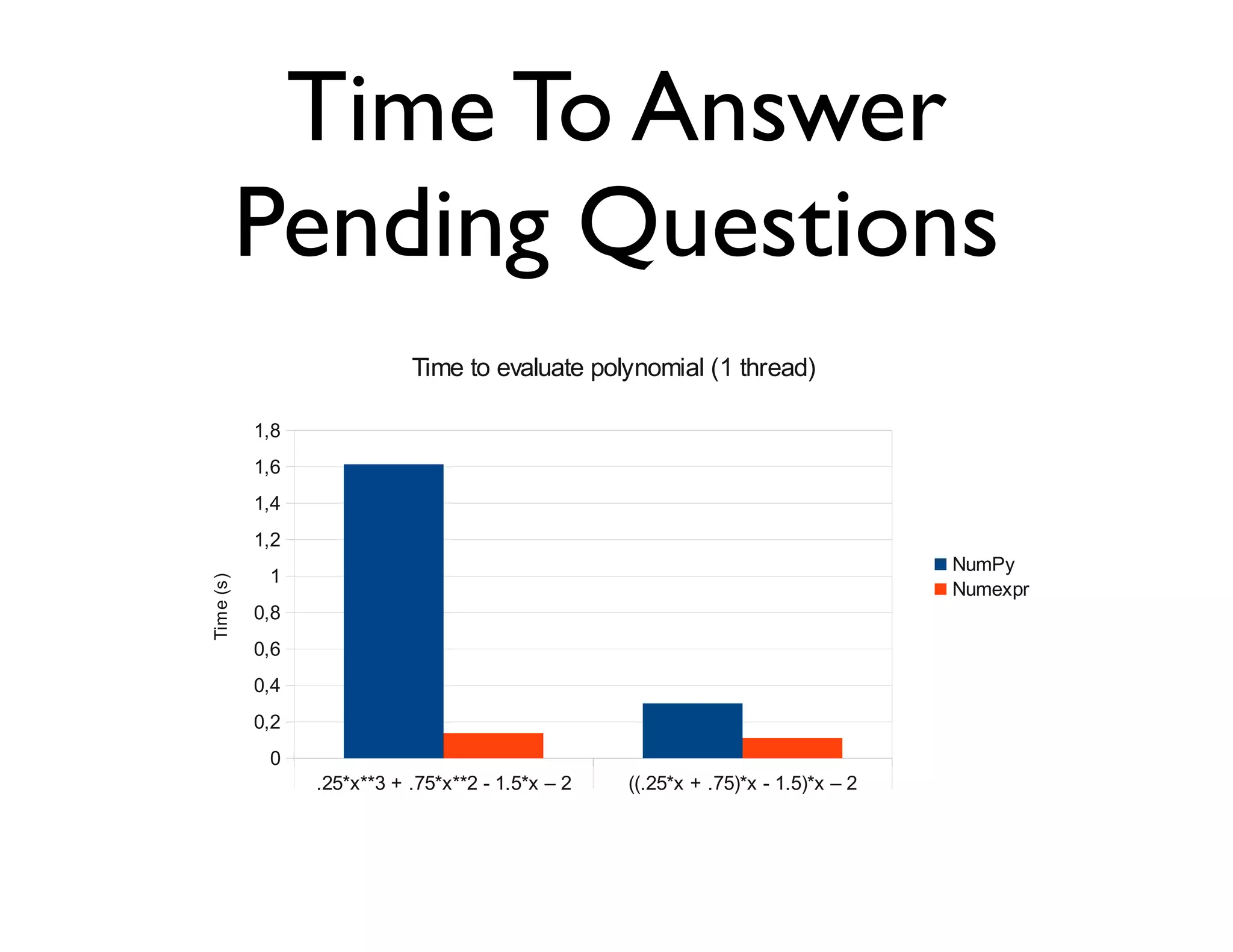
![Exercise (I)
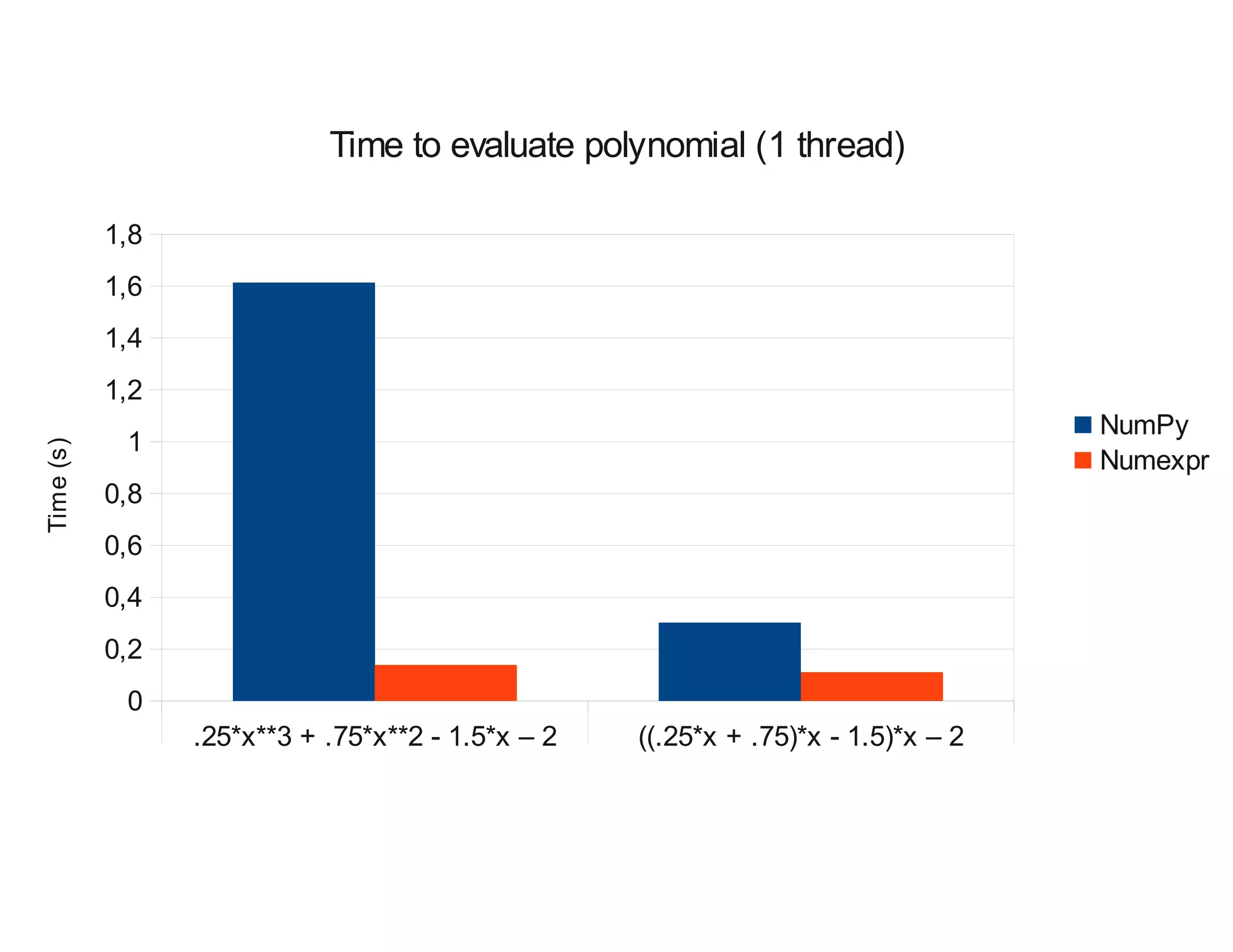
Evaluate the next polynomial:
0.25x3 + 0.75x2 + 1.5x - 2
in the range [-1, 1] with a step size of 2*10-7,
using both NumPy and numexpr.
Note: use a single processor for numexpr
numexpr.set_num_threads(1)](https://image.slidesharecdn.com/francescalted-howilearnedtostopworryingaboutcpuspeed-121128024051-phpapp01/75/Memory-efficient-applications-FRANCESC-ALTED-at-Big-Data-Spain-2012-13-2048.jpg)









![Numexpr Limitations
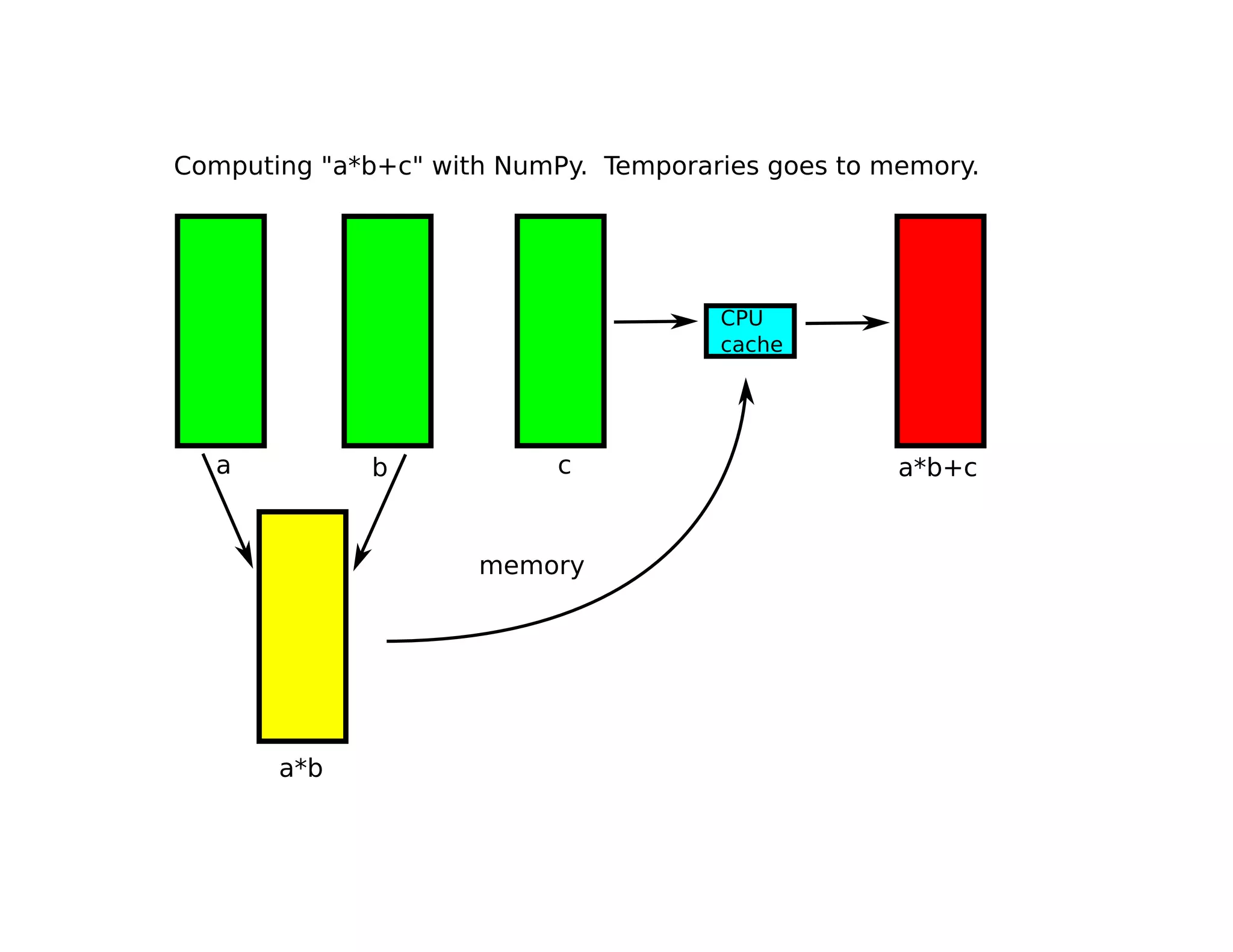
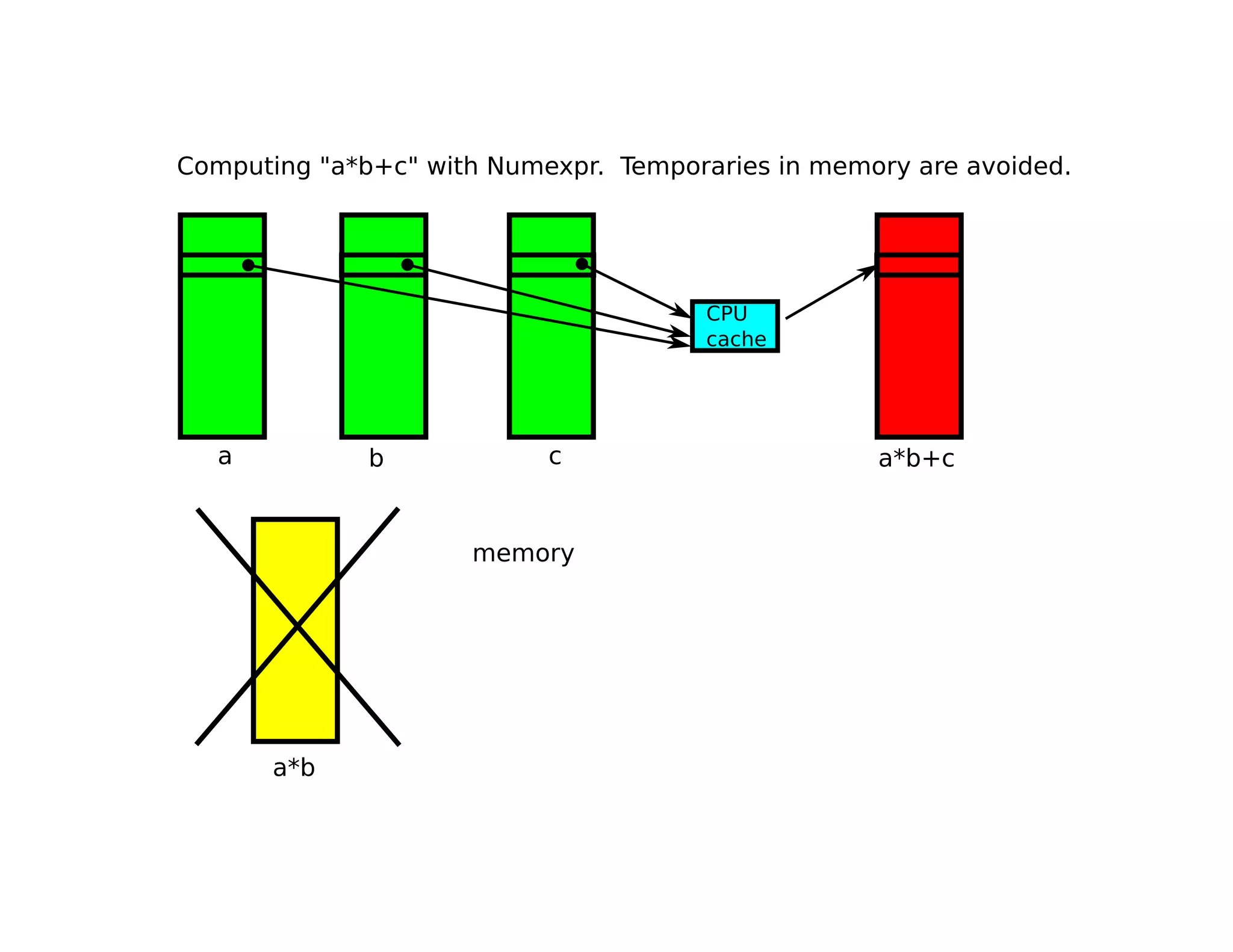
• Numexpr only implements element-wise
operations, i.e. ‘a*b’ is evaluated as:
for i in range(N):
c[i] = a[i] * b[i]
• In particular, it cannot deal with things like:
for i in range(N):
c[i] = a[i-1] + a[i] * b[i]](https://image.slidesharecdn.com/francescalted-howilearnedtostopworryingaboutcpuspeed-121128024051-phpapp01/75/Memory-efficient-applications-FRANCESC-ALTED-at-Big-Data-Spain-2012-30-2048.jpg)

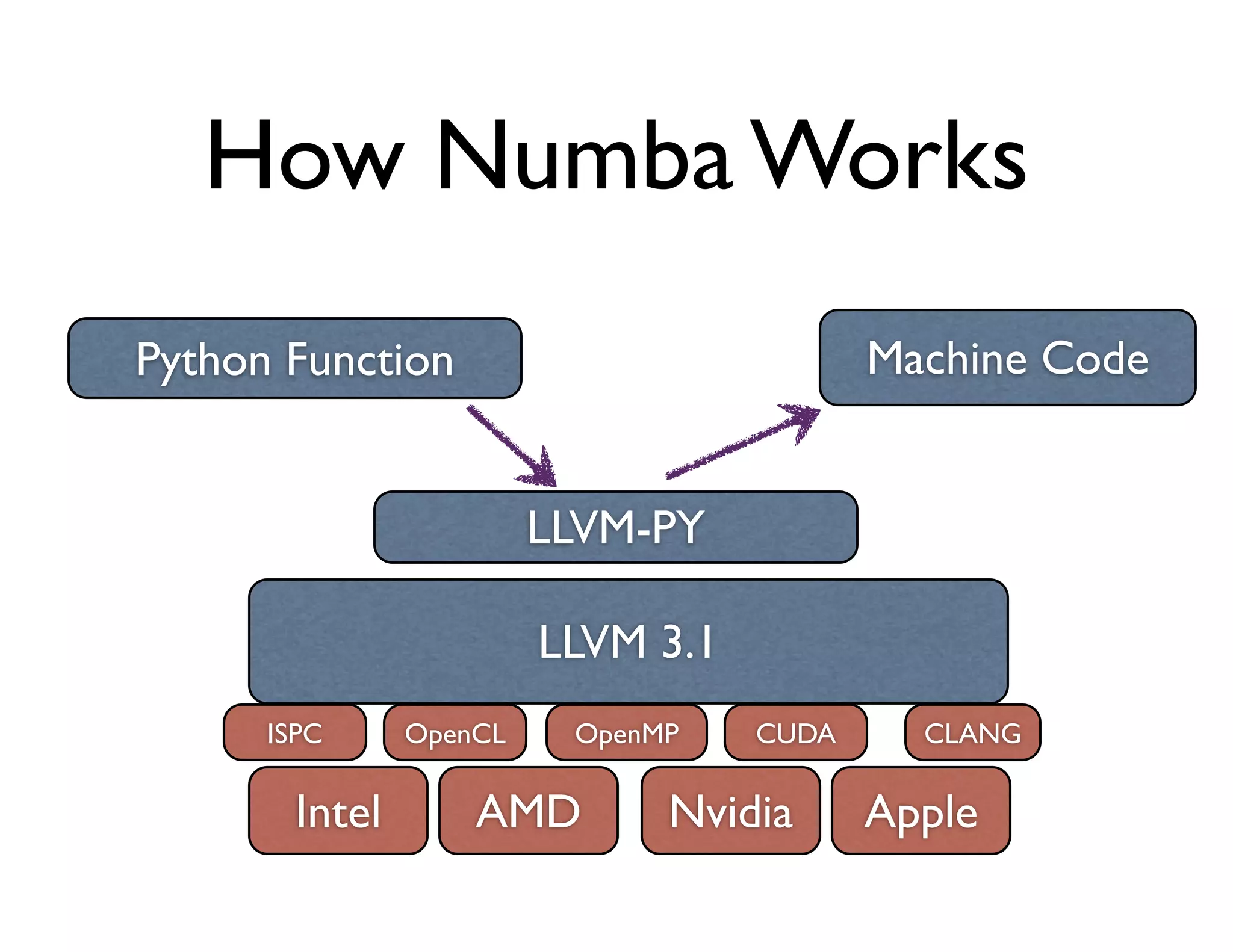
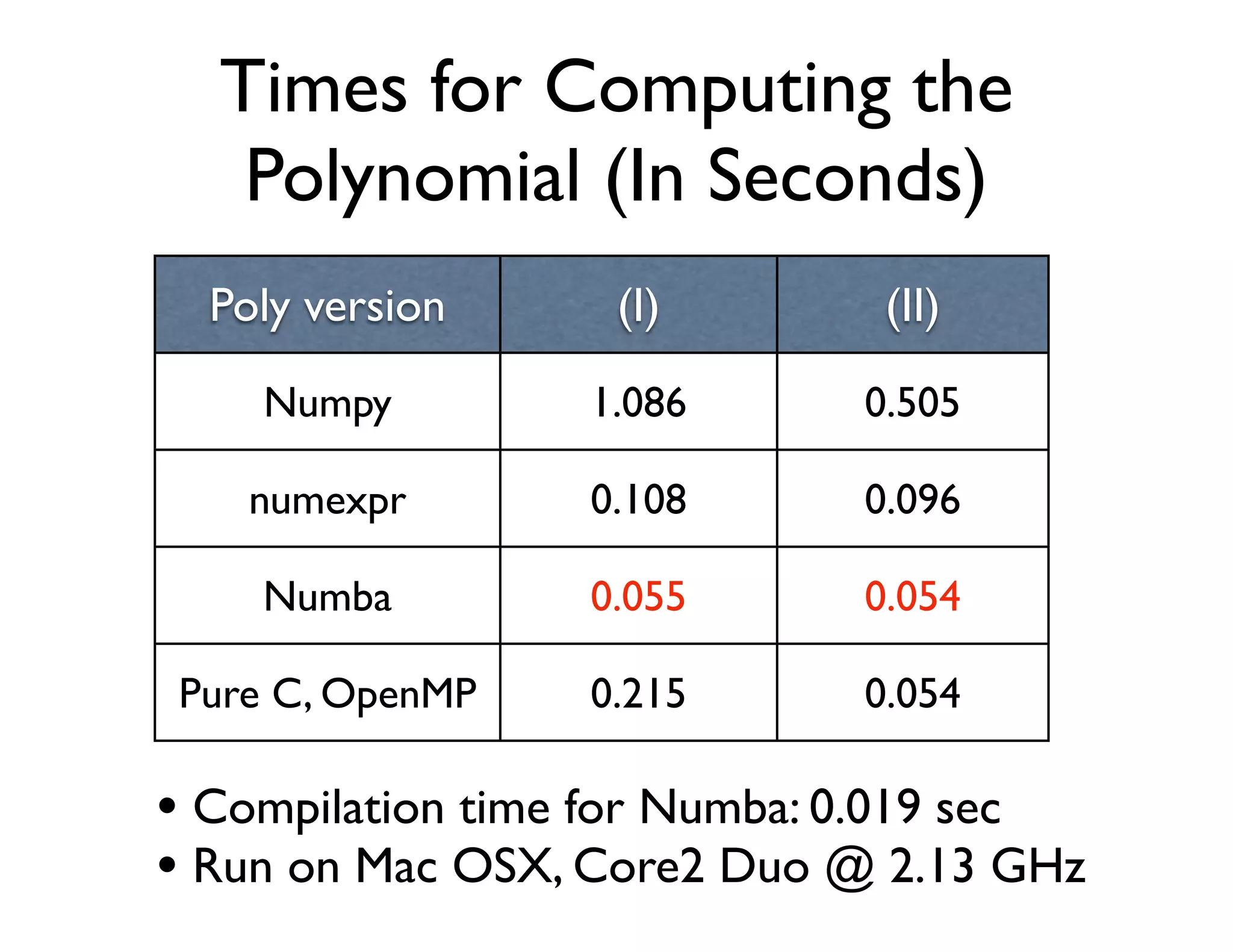
![Numba Example:
Computing the Polynomial
import numpy as np
import numba as nb
N = 10*1000*1000
x = np.linspace(-1, 1, N)
y = np.empty(N, dtype=np.float64)
@nb.jit(arg_types=[nb.f8[:], nb.f8[:]])
def poly(x, y):
for i in range(N):
# y[i] = 0.25*x[i]**3 + 0.75*x[i]**2 + 1.5*x[i] - 2
y[i] = ((0.25*x[i] + 0.75)*x[i] + 1.5)*x[i] - 2
poly(x, y) # run through Numba!](https://image.slidesharecdn.com/francescalted-howilearnedtostopworryingaboutcpuspeed-121128024051-phpapp01/75/Memory-efficient-applications-FRANCESC-ALTED-at-Big-Data-Spain-2012-33-2048.jpg)