Assignment on Recombinant DNA Technology and Gene Therapy
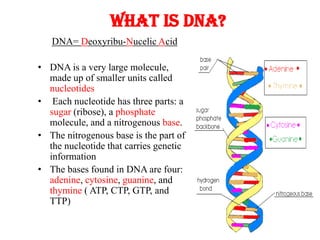
- 1. DNA= Deoxyribu-Nucelic Acid • DNA is a very large molecule, made up of smaller units called nucleotides • Each nucleotide has three parts: a sugar (ribose), a phosphate molecule, and a nitrogenous base. • The nitrogenous base is the part of the nucleotide that carries genetic information • The bases found in DNA are four: adenine, cytosine, guanine, and thymine ( ATP, CTP, GTP, and TTP) What is DNA?
- 2. • Recombinant DNA technology procedures by which DNA from different species can be isolated, cut and spliced together -- new "recombinant " molecules are then multiplied in quantity in populations of rapidly dividing cells (e.g. bacteria, yeast). Recombinant DNA Technology
- 3. • In the early 1970s it became possible to isolate a specific piece of DNA out of the millions of base pairs in a typical genome. recombinant dna technology
- 4. Recombinant DNA technology is based on a number of important things: Bacteria contain extra chromosomal molecules of DNA called plasmids which are circular. Recombinant DNA Technology
- 5. Bacteria also produce enzymes called restriction endonucleases that cut DNA molecules at specific places into many smaller fragments called restriction fragments. There are many different kinds of restriction endonucleases Recombinant DNA Technology
- 6. Restriction Enzymes and plasmid • Sticky end and blunt end are the two possible configurations resulting from the breaking of double-stranded DNA Recombinant DNA Technology
- 7. Restriction Enzymes and plasmid When RES acts at the center of symmetry, two complementary strands of DNA are of equal length, hence forms the blunt end. T C A G A T C A GA A G T C T A G T CT Recombinant DNA Technology
- 8. Restriction Enzymes and plasmid • Some RES breaks the DNA on either side of center of symmetry with the liberation of unequal fragments which are called as stick ends/ cohesive ends. • G A A T T C G A A T T C • C T T A A G C T T A A G Recombinant DNA Technology
- 9. Digestion of DNA by EcoRI to produce cohesive ends. Recombinant DNA Technology
- 10. Restriction Enzymes and plasmid • Restriction Enzymes are primarily found in bacteria and are given abbreviations based on genus and species of the bacteria. • One of the first restriction enzymes to be isolated was from EcoRI • EcoRI is so named because it was isolated from Escherichia coli strain called RY13. Recombinant DNA Technology
- 11. • A gene is a stretch of DNA that codes for a type of protein that has a function in the organism. • It is a unit of heredity in a living organism.. All living things depend on genes • Genes hold the information to build and maintain an organism's cells and pass genetic traits to offspring. What is gene?
- 12. • It can be defined as the isolation and amplification of an individual gene sequence by insertion of that individual gene sequence into a bacterium where it can be replicated Gene cloning
- 13. Step 1 A fragment of DNA, containing the gene to be cloned, is inserted into a circular DNA molecule called a vector, to produce a chimera or recombinant DNA (rDNA) molecule. 13 BASIC STEPS IN GENE CLONING
- 14. Step 2 The vector acts as a vehicle that transports the gene into a host cell, which is usually a bacterium although other types of living cell can be used. This process is called transformation.
- 15. Step 3 Within the host cell the vector multiplies producing numerous identical copies not only of itself but also of the gene that it carries.
- 16. 16 Step 4 When the host cell divides, copies of rDNA molecule are passed to the progeny and further vector replication takes place.
- 17. Step 5 After large no: of cell divisions a colony or clone of identical host cells is produced. Each cell in the clone contains one or more copies of the rDNA molecule Step 6 Then, the host cells are then lysed and rDNA can be separated.
- 18. Recombinant DNA technology had made it possible to treat different diseases by inserting new genes in place of damaged and diseased genes in the human body. Applications of rdna technology in medicine Insulin is a hormone made up of protein. It is secreted in the pancreas by some cells called as islet cells. If a person has decreased amount of insulin in his body, he will suffer from a disease called diabetes. Recombinant DNA technology has allowed the scientists to develop human insulin by using the bacteria as a host cell and it is also available in the market. It is believed that the drugs produced through microbes are safer. Insulin:-
- 19. VACCINES: Recombinant DNA technology enables the scientists to develop vaccines by cloning the gene used for protective antigen protein. Viral vaccines are most commonly developed through this technology for example, Herpes, Influenza, Hepatitis and Foot and Mouth Diseases Human Growth Hormones:- In recent years, scientists have developed many growth hormones using recombinant DNA technology. The disease of dwarfism is treated with this hormone.
- 20. Infectious Diseases:- Many diseases are diagnosed by conducting certain tests. Recombinant DNA technology has allowed the development of many tests which are being used to diagnose diseases like TB and cancer. In the diagnosis process, certain pathogens are isolated and identified, and then diagnostic kits are produced when the genome of the specific pathogen is known to kill it or block its pathogenic activity.
- 21. PRODUCTION OF NOVEL PLANTS: Rdna is used in distinguishing of novel agricultural plants which are high yielding and pest resistant Cloning of genes from wild pest resistant varieties has been used. Strain improvement for fermentation: Rdna uses extensively for improvement of strains of microbes.
- 22. REFERENCES: FUNDAMENTALS OF MEDICAL BIOTECHNOLOGY: Author: Aparna Raja Gopalan,editors: irfan ali khan, page no:203-226 U.Sathyanarayana: biotechnology: page no: 530-542 pharmaceutical biotechnology: fundamentals and applications: Author: s s kori. Page no:74-80.
- 25. Overview Definition.• Brief Description.• Why named so?• Nomenclature.• Characteristics.• Mode of Actions.• Types.• Impacts & Uses.• Use of Restriction Enzymes in• Recombinant DNA Technonoly. Restriction Enzyme Recognition• Sequences. Summary.•
- 26. RESTRICTION ENZYME/RESTRICTION ENDONUCLEASEEnzymes• that cut DNA at or near specific recognition nucleotide sequences known as restriction sites. • Especial class of enzymes that cleave (cut) DNA at a specific unique internal location along its length. • Often called restriction endonucleases (Because they cut within the molecule). Discovered• in the late 1970s by Werner Arber, Hamilton Smith, and Daniel Nathans. Essential• tools for recombinant DNA technology. Naturally• produced by bacteria that use them as a defense mechanism against viral infection. • Chop up the viral nucleic acids and protect a bacterial cell by hydrolyzing phage DNA.
- 27. RESTRICTIO N ENZYME • The bacterial DNA is protected from digestion because the cell methylates (adds methyl groups to) some of the cytosines in its DNA. • The purified forms of these bacterial enzymes are used in today's laboratories. • Commonly classified into three types, which differ in their structure and whether they cut their DNA substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. • To cut DNA, all restriction enzymes make two incisions, once through each sugar- phosphate backbone (i.e. each strand) of the DNA double helix.
- 29. Nomenclatur e• Since their discovery in the 1970s, many restriction enzymes have been identified; for example, more than 3500 different Type II restriction enzymes have been characterized. • Each enzyme is named after the bacterium from which it was isolated, using a naming system based on bacterial genus, species and strain. • For example, the name of the EcoRI restriction enzyme was derived as shown in the box. Deriva tion of the EcoRI na me Abbreviation Meaning Description E Escherichia genus co coli specific epithet R RY13 strain I First identified order of identification in the bacterium
- 31. Mode of action (how R.E. cuts DNA) The• enzyme makes two incisions, one through each of the sugar- phosphate backbones (i.e., each strand) of the double helix without damaging the nitrogenous bases. Restriction• enzymes hydrolyze the backbone of DNA between deoxyribose and phosphate groups. This leaves a phosphate group on the 5' ends and a hydroxyl on the 3' ends of both strands. A few restriction enzymes will cleave single stranded DNA, although usually at low efficiency. The• restriction enzymes most used in molecular biology labs cut within their recognition sites and generate one of three different types of ends. In the diagrams below, the recognition site is boxed in yellow and the cut sites indicated by red triangles. 5' overhangs: The enzyme cuts asymmetrically within the recognition site such that a short single-stranded segment extends from the 5' ends. BamHI cuts in this manner.
- 32. Mode of action (how R.E. cuts DNA) Blunts : Enzymes that cut at precisely opposite sites in the two strands of DNA generate blunt ends without overhangs. Smai is an example of an enzyme that generates blunt ends. The 5' or 3' overhangs generated by enzymes that cut asymmetrically are called sticky ends or cohesive ends, because they will readily stick or anneal with their partner by base pairing. The sticky end is also called a 3' overhangs: Again, we see asymmetrical cutting within the recognition site, but the result is a single-stranded overhang from the two 3' ends. KpnI cuts in this manner.
- 34. Star Activity of Restriction EnzymesStar activity is defined as the alteration in the digestion specificity that occurs under sub-optimal enzyme conditions. Star activity results in cleavage of DNA at non-specific sites. Some of the sub-optimal conditions that result in star activity are as follows: • pH >8.0 • glycerol concentration of >5% • enzyme concentration >100 units/mg of DNA • increased incubation time with the enzyme • presence of organic solvents in the reaction mixture
- 35. Artificial restriction enzymes Artificial restriction enzymes can be generated by fusing a natural or engineered DNA binding domain to a nuclease domain (often the cleavage domain of the type IIS restriction enzyme FokI). Such artificial restriction enzymes can target large DNA sites (up to 36 bp) and can be engineered to bind to desired DNA sequences. Zinc finger nucleases - are the most commonly used artificial restriction enzymes and are generally used in genetic engineering applications, but can also be used for more standard gene cloning applications.
- 36. RECOGNITION SEQUENCES AND CUTTING SITES FOR SOME ENZYMES:
- 37. Restriction Enzyme Recognition Sequences • The length of restriction recognition sites varies: The enzymes EcoRI, SacI and SstI each recognize a 6 base-pair (bp) sequence of DNA, whereas NotI recognizes a sequence 8 bp in length, and the recognition site for Sau3AI is only 4 bp in length. Length of the recognition sequence dictates how frequently the enzyme will cut in a random sequence of DNA. Enzymes with a 6 bp recognition site will cut, on average, every 46 or 4096 bp; a 4 bp recognition site will occur roughly every 256 bp.
- 38. Isoschizomer: • These are pairs of restriction enzymes specific to the same recognition sequence. • For example, SphI (CGTAC/G) and BbuI (CGTAC/G) are isoschizomers of each other. • The first enzyme discovered which recognizes a given sequence is known as the prototype e.g. SphI CUTS ----------C G T A C | G-------- BbuI CUTS ----------C G T A C | G-------- e.g. MboI CUTS ----------| G A T C -------- Sau3AI CUTS ----------| G A T C --------
- 39. Neoschizomers: • An enzyme that recognizes the same sequence but cuts it differently e.g, Smal (CCC/GGG) and XmaI (C/CCGGG) are neoschizomers of each other. AatII (recognition sequence: GACGT↓C) and ZraI (recognition sequence: GAC↓GTC) are neoschizomers of one another e.g. SmaI CUTS ----------C C C | G G G-------- XmaI CUTS --------C | C C G G G--------
- 40. Isocaudomers: • An enzyme that recognizes a slightly different sequence, but produces the same ends. i.e blunt ends e.g. BamHI CUTS ----------G | G A T C C-------- BclI CUTS ----------T | G A T C A--------
- 42. Restriction Enzyme Recognition Sequences • Restriction recognitions sites can be unambiguous or ambiguous: The enzyme BamHI recognizes the sequence GGATCC and no others - this is what is meant by unambiguous. In contrast, HinfI recognizes a 5 bp sequence starting with GA, ending in TC, and having any base between (in the table, "N" stands for any nucleotide) - HinfI has an ambiguous recognition site. XhoII also has an ambiguous recognition site: Py stands for pyrimidine (T or C) and Pu for purine (A or G), so XhoII will recognize and cut sequences of AGATCT, AGATCC, GGATCT and GGATCC. • Other point to notice from the table above is that most recognition sequences are palindromes - they read the same forward (5' to 3' on the top strand) and backward (5' to 3' on the bottom strand). Most, but certainly not all recognition sites for commonly-used restriction enzymes are palindromes.
- 43. THE IMPACT OF RESTRICTION ENZYMES Genetic engineering Type• II enzymes yielded many practical benefits, as E. K12, its genes and its vectors became the workhorses of molecular biology in the 1970s for cloning, generation of libraries, DNA sequencing, detection and overproduction of enzymes, hormones, etc.Production of insulin from recombinant bacteria and yeast by Gene technology, thus greatly increasing the supply for diabetics and the production of a recombinant vaccine for Hepatitis B by Biogen to treat the hundreds of millions of people at risk of infection by this virus
- 44. THE IMPACT OF RESTRICTION ENZYMES DNA fingerprinting • DNA fingerprinting allows the solution of paternity cases, the identification of criminals and their victims and the exoneration of the falsely accused. The use of REases in this system enabled the creation of suitable procedures for such identification. • Useful for identifying pathogenic bacterial strains, most recently of S. aureus sp with antibiotic-resistance and virulence factors mediated by mobile genetic elements, e.g. the methicillin-resistant S. aureus (MRSA) bacteria.
- 45. Why restriction enzymes are important for rDNA techniques? • Restriction enzyme recognizes and cuts, or digests, only one particular sequence of nucleotide bases in DNA • Typical restriction enzymes used in cloning experiments recognize four-, six-, or eight-base sequences • It cuts this sequence in the same way each time.. • Hundreds of restriction enzymes are known, each producing DNA fragments with characteristic ends.
- 46. GENE CLONING: DNA fragments from different species can be ligated to create Recombinant DNA
- 47. The role of a restriction enzyme in making recombinant DNA
- 48. RFLP: Restriction Fragment Length Polymorphism is a tool to study variations among individuals & among species
- 49. DNA MAPPING: Restriction Enzymes can be used to generate a restriction map. This can provide useful information in characterizing a DNA molecule.
- 50. Conclusion
- 51. Reference • https://en.wikipedia.org/wiki/Restriction_enzyme • Lehninger’s Principles of Biochemistry • http://www.biology-pages.info/R/RestrictionEnzymes.html • https://www.neb.com/products/restriction-endonucleases/restriction- endonucleases/types-of-restriction-endonucleases • http://www.bio.miami.edu/dana/dox/restrictionenzymes.html • http://www.bio.miami.edu/dana/dox/restrictionenzymes.html • https://www.dnalc.org/resources/animations/restriction.html • http://www.vivo.colostate.edu/hbooks/genetics/biotech/enzymes/renzymes.h tml • http://www.biologydiscussion.com/dna/restriction-enzymes/restriction- enzymes-in-dna-mode-of-action-and-its-types-2/12065 • https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3874209/ • http://www.sigmaaldrich.com/technical- documents/articles/biology/restriction-enzymes.html
- 54. CLONING VECTOR
- 55. INTRODUCTION A cloning vector is a small piece of DNA, taken from a virus, a plasmid, or the cell of a higher organism, that can be stably maintained in an organism, and into which a foreign DNA fragment can be inserted for cloning purposes.
- 56. FEATURES OF A CLONING VECTOR Origin of replication. Cloning site. Selectable marker. Reporter gene.
- 57. TYPES OF CLONING VECTORS 1. Plasmid. 2. Bacteriophage. 3. Cosmid. 4. Bacterial Artificial Chromosome. 5. Yeast Artificial Chromosome.
- 58. PLASMID VECTOR Plasmid vector is a small, piece of circular DNA found outside of the bacterial chromosome. Capable of replication. Can transfer autonomous genes from one cell to other.
- 59. Contains an origin of replication, allowing for replication independent of host’s genome. Contains Selective markers for the selection of cells containing a plasmid. Contains a Multiple Cloning Site (MCS). Easy to be isolated from the host cell. E.g pBR322, pUC19.
- 60. Plasmid Vectors
- 61. Properties Of Plasmid Vectors Sm aller plasmid vectors are preferred for many reasons: The efficiency of transformation is inversely related to the size of the plasmid. Lar ger plasmids are more difficult to characterize by restriction mapping.
- 62. The yield of foreign DNA is reduced with larger plasmids because these plasmids replicate to lower copy numbers
- 64. Varieties Of Plasmids Based On Functions Fertility F-plasmids which contain tra genes. They are capable of conjugation and result in the expression of sex pilli.
- 65. Resistance (R) plasmids which contain genes that provide resistance against antibiotics or poisons.
- 66. Col plasmids which contain genes that code for bacteriocins, proteins that can kill other bacteria. Degradative plasmids which enable the e.g.digestion of unusual substances, toluene and salicylic acid. Virulence plasmids which turn the bacterium into a pathogen.
- 67. Uses Of Plasmid Major use of plasmids is to make large amounts of proteins. Plasmid may also be used for gene transfer into human cells as potential treatment in gene therapy so that it may express the protein that is lacking in the cells.
- 68. BACTERIOPHAGE VECTOR Cloning Vector that uses a Bacteriophage as a means for making and storing exact copies of segments of DNA. It infects bacteria.
- 69. TYPES The bacteriophages used for cloning are ; i. phage λ ii. M13 phage.
- 70. Phage λ Vector Infect E.coli. Origin of replication. Size is 48,502 bp. High transformation efficiency, about 1000 times more efficient than the plasmid vector. Enterobacteria is important in the study of specialized transduction.
- 71. Types of Phage λ Vector a. Insertion vectors: contain a unique cleavage site whereby foreign DNA with size of 5–11 kb may be inserted. e.g λgt10 and λZAPII. b. Replacement vectors: replacement vector has two recognition sites for the restriction endonuclease used for cloning. e.g λWES.λB' and λEMBL4
- 72. DNA Cloning Using Phages As Vectors
- 73. Phage M13 Vector A gene for the lac repressor. The operator region of the lac Z gene. A lac promoter upstream of the lac Z gene. A polylinker region.
- 74. Types of M13 vector M13 mp1 form by introduction of the lacZ' genes into the m13 vector. M13 mp2 has a slightly altered lacZ' gene.It is the simplest M13 cloning vector. M13 mp7 form by the introduction of additional restriction sites into the lacZ' gene. The polylinker is inserted into the EcoRI site of M13mp2, to give M13mp7.
- 75. Complex M13 vectors have more complex polylinkers inserted into the lacz' gene, ability to take DNA fragments with two different sticky ends. M13mp9 having same polylinker as in m13 mp8 but in the reverse orientation which is important in DNA sequencing.
- 76. Use Of M13 Vector M13 vector use in nanostructures and nanotechnology.
- 77. COSMID VECTOR Hybrids between a phage DNA molecule and a bacterial plasmid. An origin of replication (ori). A cos site . An ampicillin resistance gene (amp).
- 78. Restriction cloning . sites for Cos mids can carry up to 50 kb of inserted DNA.
- 79. Uses Of Cosmids Cosmids can be used to build libraries. genomic They are often used as a cloning vector in genetic engineering.
- 80. Bacterial Artificial Chromosome DN A construct, based on a functional F-plasmid. F-plasmids play a crucial role because they contain partition genes that promote the even distribution of plasmids after bacterial cell division. Ba cterial artificial chromosome's usual insert size is 150-350 kbp.
- 81. Like other vectors, BACs contain: 1. Origin (ori) sequence derived from an E. coli plasmid called the F factor. 2. Multiple cloning sites (restriction sites). 3. Selectable markers (antibiotic resistance)
- 82. Uses Of BAC Useful for sequencing large stretches of chromosomal DNA. Frequently projects. used in genome sequencing
- 83. YEAST ARTIFICIAL CHROMOSOME YAC are genetically engineered chromosomes derived from the DNA of the yeast. Capable of carrying inserts of 100 - 1000 kbp. A YAC can be considered as a functional artificial chromosome since it includes three specific DNA sequences:
- 84. TEL: Telomere located at each chromosome end, protects the linear DNA from degradation by nucleases. CEN: Centromere which is the attachment site for mitotic spindle fibers, "pulls" one copy of each duplicated chromosome into each new daughter cell. ORI: Replication origin sequences which are specific DNA sequences.
- 85. It also contains few other specific sequences like: a. Selectable markers: that allow the easy • isolation of yeast cells that have taken up the artificial chromosome. b. Recognition site: for the two restriction • enzymes EcoRI and BamHI.
- 87. Uses Of YAC Used to express eukaryotic proteins that require posttranslational modification. Us ed for detailed mapping of specific regions of the genome.
- 88. REFERENCES • 1. Hall, RM; Collis, CM (1995). "Mobile gene cassettes and integrons: Capture and spread of genes by site-specific recombination". Molecular microbiology • 2. Plant biotechnology – the genetic manipulatio by Adrian Slater, Nigel W.Scott & Mark R. Fowlerns of plants 2nd edition • • 3. Principles of gene Manipulations & Genomics – Primrose • 4. Joseph Sambrook, David Russell. "Chapter 1". Molecular Cloning • Laboratory Manual 1 (3rd ed.) • 5. Lederberg, J and Lederberg, EM (1952) Replica plating and indirect selection of bacterial mutants. J Bacteriol.
- 91. • Submitted To: • DR. GOVIND SINGH ASSISTANT PROFESSOR • PHARMACOLOGY • Submitted By: • KAPIL YADAV M.PHARM PHARMACOLOGY•
- 92. Recombinant DNA Technology • Revolutionized biology • Manipulation of DNAsequences and the construction of chimeric molecules, provides a means of studying how a specific segment of DNAworks • Studies in bacteria and bacterial viruses have led to methods to manipulate and recombine DNA • Once properly identified, the recombinant DNAmolecules can be used in various ways useful in medicine and human biology
- 95. Recombinant Pharmaceuticals • Anumber of human disorders can be traced to the absence or malfunction of a protein normally synthesized in the body. • Most of these disorders can be treated by supplying the patient with the correct version of the protein • Hence, modern pharmaceutical manufacturing frequently relies upon recombinant drugs.
- 96. Recombinant Pharmaceuticals Hu• man Insulin Human• Growth Hormone Human• blood clotting factors • Vaccines Mono• clonal Antibodies Interfe• rons Anti• biotics & other secondary metabolites
- 97. Human Insulin Earl• iest use of recombinant technology Modify• E.coli cells to produce insulin; performed by Genentech in 1978 Prio• r, bovine and porcine insulin used but induced immunogenic reactions Also, the• re were many purification and contamination hassles. • Toovercometheseproblems,researchersinserted human insulin genes into a suitable vector (E.coli)
- 98. • Insulin is a hormone made up of protein. It is secreted in the pancreas by some cells called as islet cells. If a person has decreased amount of insulin in his body, he will suffer from a disease called diabetes. Recombinant DNA technology has allowed the scientists to develop human insulin by using the bacteria as a host cell and it is also available in the market. It is believed that the drugs produced through microbes are safer. Applications of rdna technology in medicine Recombinant DNA technology had made it possible to treat different diseases by inserting new genes in place of damaged and diseased genes in the human body.
- 99. Producing Recombinant Insulin • First, scientists synthesized genes for the two insulinA& B chains. • They were then inserted into plasmids along with a strong lacZ promoter. • The genes were inserted in such a way that the insulin & B-galactosidase residues would be separated by a methionine residue. This is so that the insulinA& B chains can be separated easily by adding cyanogen bromide.
- 100. Producing Recombinant Insulin The• vector was then transformed into E.coli cells. Once• inside the bacteria, the genes were "switched-on" by the bacteria to translate the code into either the "A" chain or the "B" chain proteins found in insulin The• purified insulin A and B chains were then attached to each other by disulphide bond formation under laboratory conditions
- 102. Human Growth Hormones Soma• tostatin and Somatotrophin are two proteins that act in conjunction to control growth processes in the human body, their malfunction leading to painful and disabling disorders such as Acromegaly (uncontrolled bone growth) and Dwarfism. Somatos• tatin was the first human protein to be synthesized in E. coli. Being a very short protein, only 14 amino acids in length, it was ideally suited for artificial gene synthesis.
- 103. Production of Recombinant Human Growth Hormones • The strategy used was the same as described for recombinant insulin, involving insertion of the artificial gene into a lacZ′ vector, synthesis of a fusion protein, and cleavage with cyanogen bromide
- 104. Recombinant Blood Clotting Factors • Human factor VIII is a protein that plays a central role in blood clotting. • The commonest form of haemophilia in humans results from an inability to synthesize factor VIII • The factor VIII gene is very large. The mRNA codes for a large polypeptide (2351 amino acids), which undergoes a complex series of post-translational processing events, eventually resulting in a dimeric protein consisting of a large subunit and a small subunit.
- 105. Production of Recombinant Human Blood Clotting Factors The• two subunits contain a total of 17 disulphide bonds and a number of glycosylated sites. As might be anticipated for such a large and complex protein, it has not been possible to synthesize an active version in E. coli. • TwoseparatefragmentsfromthecDNAwereused. Each cDNA fragment was ligated into an expression vector along with Ag promoter (a hybrid between the chicken b-actin and rabbit b-globin sequences) and a polyadenylation signal from SV40 virus. The pla• smid was introduced into a hamster cell line and recombinant protein obtained.
- 106. Production of Recombinant Human Blood Clotting Factors Alter• native method- pharming The comp• lete human cDNA has been attached to the promoter for the whey acidic protein gene of pig, leading to synthesis of human factor VIII in pig mammary tissue and subsequent secretion of the protein in the milk. The• factor VIII produced in this way appears to be exactly the same as the native protein
- 107. Recombinant Vaccines • Two types: (i)Recombinant protein vaccines: This is based on production of recombinant DNAwhich is expressed to release the specific protein used in vaccine preparation (ii)DNAvaccines: Here the gene encoding for immunogenic protein is isolated and used to produce recombinant DNAwhich acts as vaccine to be injected into the individual.
- 108. Recombinant protein vaccines: • Apathogen produces its proteins in the body which elicit an immune response from the infected body. • The gene encoding such a protein is isolated from the causative organism • This DNAis expressed in another host organism, like genetically engineered microbes; animal cells; plant cells; insect larvae etc, resulting in the release of appropriate proteins. • These when injected into the body, causes immunogenic response against the corresponding disease providing immunity.
- 109. DNAvaccines: Refers to• the recombinant vaccines in which the DNA is used as a vaccine. The• gene responsible for the immunogenic protein is identified, isolated and cloned with corresponding expression vector. Upon int• roduction into the individuals to be immunized, it produces a recombinant DNA. This• DNA when expressed triggers an immune response and the person becomes successfully vaccinated. The• mode of delivery of DNA vaccines include: direct injection into muscle; use of vectors like adenovirus, retrovirus etc; invitro transfer of the gene into autologous cells and reimplantation of the same and particle gun delivery of the DNA.
- 110. DNA vaccines: In• certain cases, the responsible gene is integrated into live vectors which are introduced into individuals as vaccines. This• is known as live recombinant vaccines. Eg: vaccinia virus. Live vaccinia virus vaccine (VV vaccine) with genes corresponding to several diseases, when introduced into the body elicit an immune response but does not actually cause the diseases.
- 111. Recombinant Antibodies • An immunoglobulin which produced because of the introduction of an antigen into the body, and which possesses the ability to recognize the antigen. • Using recombinant antibody has significant advantages compared with the conventional antibody and there for its use becoming more popular now days. • The fact that no animals are needed in the manufacturing procedure of the recombinant antibodies, in addition, the manufacturing time is relatively short compared with the conventional method. • Moreover, the quality of the final product is higher
- 112. Production of Recombinant Antibodies • The production of non-animal recombinant antibodies can be broken down into five steps: (1)creation of an antibody gene library (2) display of the library on phage coats or cell surfaces (3)isolation of antibodies against an antigen of interest (4) modification of the isolated antibodies and (5)scaled up production of selected antibodies in a cell culture expression system.
- 113. Interferons Interfe• rons (IFNs) are a group of signalling proteins made and released by host cells in response to the presence of pathogens, such as viruses, bacteria, parasites, or tumor cells. In a typical• scenario, a virus-infected cell will release interferons causing nearby cells to heighten their anti-viral defenses.
- 114. Production of Recombinant Interferons Recomb• inant DNA technology has proved the most satisfactory route to the large scale production of human interferons. The g• enes of all three types of HuIFN have been cloned in micro-organisms and expression obtained. Hu• IFNȕ and Ȗproduced in this manner lack the glycosylation present in the naturally occurring substances but this does not affect their specific activity.
- 115. Production of Recombinant Interferons • Greatly improved methods of purification, including immuno-adsorption chromatography on monoclonal antibody columns, are now available so there should be no difficulty in supplying adequate amounts of very pure interferon of all three types although, up till now, only HuIFNα has been readily available.
- 116. Recombinant Secondary Metabolites • The importance of antibiotics to medicine has led to much research into their discovery and production. • GM micro-organisms are used to increase • production. • Another technique used to increase yields is gene amplification, where copies of genes • coding for enzymes involved in the antibiotic • production can be inserted back into a cell, via vectors such as plasmids.
- 117. Production of Recombinant Plant Secondary Metabolites Plant• secondary metabolites can also be produced by rDNA technology in plant suspension cultures, micro-organism cultures and hairy root cultures • A. rhizogenes mediated transformation which can transfer foreign genes into the transformed hairy root. • E.g.: 6-hydroxylase gene of Hyoscyamus muticus which was introduced to Atropa belladonna using A. rhizogenes. Eng• ineered roots showed an increased amount of enzyme activity and a five-fold higher concentration of scopolamine.
- 118. REFERENCE • 1. Hall, RM; Collis, CM (1995). "Mobile gene cassettes and integrons: Capture and spread of genes by site-specific recombination". Molecular microbiology • 2. Plant biotechnology – the genetic manipulatio by Adrian Slater, Nigel W.Scott & Mark R. Fowlerns of plants 2nd edition • • 3. Principles of gene Manipulations & Genomics – Primrose • 4. Joseph Sambrook, David Russell. "Chapter 1". Molecular Cloning Lab• oratory Manual 1 (3rd ed.) • 5. Lederberg, J and Lederberg, EM (1952) Replica plating and indirect selection of bacterial mutants. J Bacteriol.
- 121. GENE THERAPY-VARIOUS TYPES OF GENE TRANSFER TECHNIQUES PRESENTED BY:-KAPIL YADAV
- 148. GENE TRANSFER • Gene transfer is defined simply as a technique to stably introduce foreign genes into the genome of target cells. • The directed desirable gene transfer from one organism to another and the subsequent stable integration & expression of foreign gene into the genome is referred as genetic transformation. • Transient transformation occur when DNA is not integreted into host genome
- 149. • Stable transformation occur when DNA is integrated into host genome and is inherited in subsequent generations. • The transferred gene is known as transgene and the organism that develop after a successful gene transfer is known as transgenic.
- 150. METHODS OF GENE TRANSFER DNA transfer by natural methods • 1. Conjugation • 2. Bacterial transformation • 3. Retroviral transduction
- 151. DNA TRANSFER BYARTIFICIAL METHODS • Physical methods • 1. Microinjection • 2. Biolistics transformation • Chemical methods • 1. DNA transfer by calcium phosphate method • 2. Liposome mediated transfer • Electrical methods • 1. Electroporation
- 152. CONJUGATION • Requires the presence of a special plasmid called the F plasmid. • Bacteria that have a F plasmid are referred to as as F+ or male. Those that do not have an F plasmid are F- of female. • The F plasmid consists of 25 genes that mostly code for production of sex pilli. • A conjugation event occurs when the male cell extends his sex pilli and one attaches to the female.
- 153. • This attached pilus is a temporary cytoplasmic bridge through which a replicating F plasmid is transferred from the male to the female. • When transfer is complete, the result is two male cells. • When the F+ plasmid is integrated within the bacterial chromosome, the cell is called an Hfr cell (high frequency of recombination cell).
- 155. TRANSFORMATION Transformation is the direct uptake of exogenous DNA from its surroundings and taken up through the cell membrane . Transforma• tion occurs naturally in some species of bacteria, but it can also be effected by artificial treatment in other species. Cel• ls that have undergone this treatment are said to be competent. Any DNA th• at is not integrated into the chromosome will be degraded.
- 157. TRANSDUCTION Gene• transfer from a donor to a recipient by way of a bacteriophag.. If• the lysogenic cycle is adopted, the phage chromosome is integrated (by covalent bonds) into the bacterial chromosome, where it can remain dormant for thousands of generation The• lytic cycle leads to the production of new phage particles which are released by lysis of the host.
- 159. VECTORLESS or DIRECT GENE TRANSFER• Physical methods • 1. Microinjection • • • • • • • 2. Biolistics transformation Chemical methods 1. DNA transfer by calcium phosphate method 2. Liposome mediated transfer 3. Transfer of DNA by use of polyethene glycol Electrical methods 1. Electroporation
- 160. Electroporation • Electroporation uses electrical pulse to produce transient pores in the plasma membrane thereby allowing DNA into the cells. • These pores are known as electropores. •
- 161. The c• ells are placed in a solution containing DNA and subjected to electrical pulse to cause holes in the membrane. The fore• ign DNA fragments enter through holes into the cytoplasm and then to nucleus.
- 162. Advantages of Electroporation • 1. Method is fast. • 2. Less costly. • 3. Applied for a number of cell types. • 4. Simultaneously a large number of cell can be treated. • 5. High percentage of stable transformants can be produced
- 163. Microinjection The microinjection is the process of transferring the desirable DNA into the living cell ,through the use of glass micropipette . Glass micropipette is usually of 0.5 to 5 micrometer, easily penetrates into the cell membrane and nuclear envelope. The desired gene is then injected into the sub cellular compartment and needle is removed
- 165. Limitations of microinjection • Costly. • Skilled personal required. • More useful for animal cells.
- 166. Biolistics or Microprojectiles Biol• istics or particle bombardment is a physical method that uses accelerated microprojectiles to deliver DNA or other molecules into intact tissues and cells. The gene gun is a device that litera• lly fires DNA into target cells . The DNA to be• transformed into the cells is coated onto • microscopic beads made of either gold or tungsten.
- 167. • The coated beads are then attached to the end of the plastic bullet and loaded into the firing chamber of the gene gun. • An explosive force fires the bullet with DNA coated beads towards the target cells that lie just beyond the end of the barrel. • Some of the beads pass through the cell wall into the cytoplasm of the target cells
- 169. Liposome mediated gene transfer • Liposomes are spheres of lipids which can be used to transport molecules into the cells. • These are artificial vesicles that can act as delivery agents for exogenous materials including transgenes. • Promote transport after fusing with the cell membrane. • Cationic lipids are those having a positive charge are used for the transfer of nucleic acid.
- 170. Advantag es • 1. Simplicity. • 2. Long term stability. • 3. Low toxicity. • 4. Protection of nucleic acid from degradation
- 171. Calcium phosphate mediated DNA transfer • The process of transfection involves the admixture of isolated DNA (10-100ug) with solution of calcium chloride and potassium phosphate so precipitate of calcium phosphate to be formed. • Cells are then incubated with precipitated DNA either in solution or in tissue culture dish. • A fraction of cells will take up the calcium phosphate DNA precipitate by endocytosis.
- 172. Transfe• ction efficiencies is quite low.
- 173. Polyethylene glycol mediated transfectionThis• method is utilized for protoplast only. Polyethylene glycol stimul• ates endocytosis and therefore DNA uptake occurs. Pr• otoplasts are kept in the solution containing polyethylene glycol (PEG). After transfer• of DNA to the protoplast in presence of PEG and other chemicals, PEG is allowed to get removed
- 174. • Sukharev, S.I., Klenchin, V.A., Serov, S.M., Chernomordik, L.V. & Chizmadzhev, Y.A. (1992). Electroporation • and electrophoretic DNA transfer into cells. The effect of DNA interaction with electropores. Biophys. J., 63 (5): • 1320-1327. • [2] Chu, G., Hayakawa, H. & Berg, P. (1987). Electroporation for the efficient transfection of mammalian cells with • DNA. Nucleic Acids Res., 15 (3): 1311-1326. • [3] Akamatsu, W., Okano, H.J., Osumi, N., Inoue, T., Nakamura, S., Sakakibara, S.I., Miura, M., Matsuo, N., Darnell, • R.B. & Okano, H. (1999). Mammalian ELAV-like neuronal RNAbinding proteins HuB and HuC promote • neuronal development in both the central and the peripheral nervous systems. Proc. Natl. Acad. Sci. USA., 96 (17): • 9885-9890. • [4] Osumi, N. & Inoue, T. (2001). Gene transfer into cultured mammalian embryos by electroporation. Methods., 24 • Antoni Ivorra, Boris Rubinsky. "Gels with predetermined conductivity used in electroporation of tissue USPTO Application #: 20080214986 - Class: 604 21 (USPTO)". • Jump upJump up^ Sugar, I.P.; Neumann, E. (1984). "Stochastic model for electric field-induced membrane pores electroporation". Biophysical Chemistry 19 (3): 211–25. doi:10.1016/0301- 4622(84)87003-9. PMID 6722274. • ^ Alberts, Bruce; et al. (2002). Molecular Biology of the Cell. New York: Garland Science. p. G:35. ISBN
- 175. • Jogdand, S.N. (2006). Gene Biotechnology. Himalaya Publishing House. Mumbai, India. 2nd ed., p 237- 249. • Chen, C.A. & Okayama, H. (1988). Calcium phosphate-mediated gene transfer: a highly efficient trasfection system for stably transforming cells with plasmid DNA. Biotechniques., • Watwe, R.M. & Bellare, J.R. (1995). Manufacture of liposomes a review. Curr. Sci. India., . Nicolau, C., Legrand, A. & Grosse, E. (1987). Liposomes as carriers for in vivo gene transfer and expression. Method Enzymol., 149: 157-176. • lies, M.A. & Balaban, A.T. (2001). Recent developments in cationic lipid-mediated gene delivery and gene therapy. Expert Opin. Ther. Patents., • Reece, R.J. (2004). Analysis of Genes and Genomes. John Wiley and Sons Ltd. • Johnston, S.A. & Tang, D.C. (1994). Gene gun transfection of animal cells and genetic immunization. Methods Cell Biol., 43 Klein, T.M., Arentzen, R., Lewis, P.A. & Fitzpatrickmcelligott, S. (1992). Transformation of microbes, plants and animals by particle bombardment. Biotechnology • King, R. (2004). Gene delivery to mammalian cells by microinjection. Methods Mol. Biol., • David B. Burr; Matthew R. Allen (11 June 2013). Basic and Applied Bone Biology. Academic. p. 157. ISBN 978- 0-12-391459-0. Retrieved 15 July 2013. • ^ Juan Carlos Lacal; Rosario Perona; James Feramisco (11 June 1999). Microinjection. Springer. p. 9. ISBN •
- 177. APPLICATIONS AND RECENT ADVANCES OF GENE THERAPY • PRESENTED BY: • Kapil Yadav • M.PHARM 1st YEAR • Pharmacology PRESENTED TO:• Dr. Govind• Singh Assistant• Professor
- 179. WHAT IS GENE THERAPY Gene therapy is an experimental technique that uses genes to• treat or prevent disease. In the future, this technique may allow doctors to treat a• disorder by inserting a gene into a patient cells instead of using drugs or surgery. Gene therapy was first conceptualized in• 1972. Researchers are testing several approaches to gene therapy,• including: Replacing a mutated gene that causes disease with a healthy copy of a gene. Inactivating, or “knocking out” a mutated gene that is functioning improperly. Introducing a new gene into the body to help fight a disease.
- 181. APPLICATIONS OF GENE THERAPY Gene therapy is used for the treatment of disorder involving blood cells can be cured by this method. The hematopoietic stem cells are removed from an affected individual and transfected with functional genes. The engineered stem cells are then re- injected into the individual. This method is used with individual suffering from SCID resulting from a defective gene encoding adenosine diaminase (ADA).
- 185. 1. THERAPY FOR CYSTIC FIBROSIS Cystic fibrosis is a fatal genetic disease characterized by accumulation of sticky ,dehydrated mucous in respiratory tract and lungs In the patient of Cystic fibrosis the CFTR (Cystic fibrosis transmembrane regulator) protein is not produced due to gene defect. The chloride ion concentrate within the cell which draw water from surrounding. As a result respiratory tract and lungs becomes dehydrated with sticky mucous.
- 187. CONTINUED… In Gene therapy, adenoviral vector system have been used. Adenovirus do not integrate themselves into host cells. By using adeno-associated virus vector system, some good results were reported in the gene therapy of cystic fibrosis.
- 192. 2. THERAPY FOR SCID (ADA DEFICIENCY) Severe Combined Immunodeficiency Disease• This is inherited immune disorder associated with T - lymphocytes, and B-lymphocytes dysfunction. Deficiency of ADA accumulate and destroy T -lymphocytes. T-lymphocytes are essential for body ̍s immunity. Patient of SCID (lacking ADA) die at an young age.
- 193. TREATMENT • A plasmid vector bearing a proviral DNA is selected. • A part of proviral DNA is replaced by ADA gene. • Circulating lymphocytes are removed. • The cells are transfected with ADA gene. • The genetically-modified lymphocytes are
- 194. Continued… • Infuse lymphocytes with ADA gene. • Expressed into patient. • Synthesis of ADA in child with SCID • Correction of SCID
- 197. 3. Therapy for Lesch-Nyhan syndrome • The most likely candidates for future gene therapy trials will be rare diseases such as Lesch-Nyhan syndrome, a distressing disease in which the patients are unable to manufacture a particular enzyme. This leads to a bizarre impulse for self- mutilation, including very severe biting of the lips and fingers. The normal version of the defective gene in this disease has now been cloned.
- 199. X-linked recessive Disease LNS• is transmitted as and X- linked recessive trait. Female carriers do not show the symptoms. LNS is characterized by self- mutilating behaviours such as lip and finger biting and/or head banging. The deficiency of HGPRT activity leads to accumulation of guanine• . By using retroviral vector• system, HGPRT producing genes were inserted into cultured human bone marrow cells.
- 201. 4. THERAPY FOR HEMOPHILIA • Hemophilia is a genetic disease due to lack of a gene that encodes for clotting factor IX. • It is characterized by excessive bleeding. • By using a retroviral system ,genes for the synthesis of factor IX were inserted into the liver cells.
- 205. 5. THERAPY FOR BLINDNESS(LCA) • Leber's congenital amaurosis (LCA) is a rare • inherited eye disease that appears at birth or in • the first few months of life. • Characterized by no pupillary responses and • severe vision loss and blindness. • Researchers at Moorfields Eye Hospital and • University College London in London conducted • the first gene therapy clinical trial for patients • with RPE65 LCA.
- 207. CHM-(CHOROIDEREMIA) Choroideremia is caused by a loss-of-function mutation in the CHM gene which encodes Rab escort protein 1 (REP1), a protein involved in lipid modification of Rab proteins While the complete mechanism of disease is not fully understood, the lack of a functional protein in the retina results in cell death and the gradual deterioration of the choroid, retinal pigment epithelium (RPE), and retinal photoreceptor cells. As of 2017, there is no treatment for choroideremia; however, retinal gene therapy clinical trials have demonstrated a
- 208. 6. THERAPY FOR CANCER • � Multiple gene therapy strategies have been developed to treat a wide variety of cancers,• including suicide gene therapy, anti• - angiogenesis and therapeutic gene vaccines.• • � Two-thirds of all gene therapy trials are for cancer and many of these are entering the• advanced stage e.g. gene vaccine trials for• prostate cancer and pancreas cancer.•
- 210. 7. THERAPY FOR HIV • FDA Approves Further Study Of Promising Gene • Therapy HIV Treatment (19 March, 2015). • Experimental stem cell gene therapy that could act as • functional cure for HIV infection has been approved by • the FDA to move into early human test trials. • Cells harvested from a positive person’s body. The stem • cells are genetically manipulated to develop into white • blood cells that are missing the key cellular receptors • that the HIV virus uses to insert its genetic code into • healthy cells. The modification effectively models a HIV positive • person’s white blood cells .
- 212. 8. THERAPY FOR GAUCHER DISEASE Gaucher disease is a lysosomal storage disease which is characterized by deficient activity of lysosomal enzyme, known as glucocerebrosidase. This resulted in progressive accumulation of glucocerebroside only in bone marrow. The extensive study based on transferring therapeutic gene to hematopoietic stem cell.
- 216. RECENT ADVANCES IN GENE THERAPY
- 217. The Beginning… • � In the 1980s, Scientists began to look into gene therapy.• • � They would insert human genes into a bacteria cell. • � Then the bacteria cell would transcribe and translate the information into a protein • � Then they would introduce the protein into human cells
- 218. • On September 14, 1990 at the U.S. National Institutes of Health, W. French • Anderson M.D. and his colleagues R. Michael Blaese, M.D., C. Bouzaid, M.D., and Kenneth Culver, M.D., performed the first approved gene therapy procedure on four-year old Ashanthi DeSilva, Born with a rare genetic disease called severe combined immunodeficiency (SCID). • What did they do • In Ashanthi's gene therapy procedure, doctors removed white blood cells from the • child's body, let the cells grow in the laboratory, inserted the missing gene into the • cells, and then infused the genetically modified blood cells back into the patient's • bloodstream. • First Approved Gene Therapy
- 220. Current Status FDA hasn’t approved any human gene therapy product for sale. Reasons: In 1999, 18-year-old Jesse Gelsinger died from multiple organ failure 4 days after treatment for ornithine transcarboxylase deficiency. • . Death was triggered by severe immune response to adenovirus carrier. January 2003, halt to using retrovirus vectors in blood stem cells because children developed leukemia-like condition after successful treatment for X-linked severe combined immunodeficiency disease.
- 221. RECENT ADVANCES IN GENE THERAPY • Although early clinical failures led many to dismiss gene therapy as • over-hyped, clinical successes since 2006 have bolstered new optimism • in the promise of gene therapy. • These include successful treatment of patients with the retinal disease, X-linked SCID, chronic lymphocytic leukemia(CLL), acute lymphocytic leukemia(ALL), multiple myeloma and Parkinson's disease. • • These recent clinical successes have led to a renewed interest in gene • therapy, with several articles in scientific and popular publications • calling for continued investment in the field.
- 223. More than 5000 patients have been treated in last ~12 years worldwide.
- 226. A success story • As of early 2007, she was still in good health, and • she was attending college. Some would state that the • study is of great importance despite its indefinite • results, if only because it demonstrated that gene • therapy could be practically attempted without • adverse consequences
- 227. Cindy Kisik and Ashanthi in 1992 with the pioneer physicians of gene therapy: (from left) French Anderson, MD; Michael Blaese, MD; and Kenneth Culver.
- 228. R. Michael Blaese, MD with Ashanthi DeSilva (left) and Cindy Kisik at the IDF 2013 National Conference, June 29
- 229. 'mending broken hearts' by using gene therapy Novel• techniques to “mend broken hearts” using gene therapy and stem cells represent a major new frontier in the treatment of heart disease• • � It was achieved by the researchers at Gladstone Institute of Cardiovascular Disease in California• • � They were able to re-programme scar-forming cells into heart muscle cells, some of which were capable of transmitting the kind of electrical signals that make the heart beat• • � They performed on a live mice, transforming scar-forming cells, called fibroblasts, into beating heart muscle cells• • � They injected three genes (cocktail of genes) into the heart of live mice that had been damaged by heart attack, fibroblasts could be turned into working heart cells.• • � Researchers said that the “cocktail of genes” used to regenerate cells could one day be replaced with• “small drug-like molecules” that would offer safer and easier delivery
- 230. First Real-Time MRI-Guided Gene Therapy for Brain Cancer • �Neurosurgeons at the University of California, San Diego School of Medicine and UC San Diego• Moores Cancer Center are among the first in the world to utilize real-time magnetic resonance imaging (MRI) guidance for delivery of gene therapy as a potential treatment for• brain tumors• • �Using MRI navigational technology, neurosurgeons can inject Toca 511 (vocimagene amiretrorepvec• ), a novel investigational gene therapy, directly into a brain malignancy • �The new approach offers a precise way to deliver a therapeutic virus designed to make the tumor susceptible to cancer• -killing drugs
- 231. Continued… • Toca 511 is a retrovirus engineered to • selectively replicate in cancer cells, such as • glioblastomas. • Toca 511 produces an enzyme that converts • an anti-fungal drug, flucytosine (5-FC), into • the anti-cancer drug 5-fluorouracil (5-FU). • After the injection of Toca 511, the patients • are treated with an investigational extended release • oral formulation of 5-FC called Toca • FC. • Cancer cell killing takes place when 5-FC • comes into contact with cells infected with • Toca 511.
- 232. stem cell gene therapy gives hope to prevent inherited neurological disease • Scientists from The University of Manchester have used stem cell gene • therapy to treat a fatal genetic brain disease • It was used to treat Sanfilippo – a fatal inherited condition which causes • progressive dementia in children • Sanfilippo, is currently untreatable mucopolysaccharide (MPS) disease • It is caused by the lack of SGSH enzyme in the body which helps to breakdown • and recycle long chain sugars, such as heparan sulphate (HS) • Children with the condition build up and store excess HS throughout their body • from birth which affects their brain and results in progressive dementia and • hyperactivity, followed by losing the ability to walk and swallow
- 233. Continued… • � Researchers have developed a stem cell gene therapy which overproduces the SGSH enzyme specifically in bone marrow white blood cells to increase• SGSH enzyme from bone marrow transplants, and to target it to the cells that• traffic into the brain• • � It was seen that mice treated by this method produce five times the normal SGSH enzyme levels in the bone marrow and• and 11 per cent of normal levels in the brain• • � The enzyme is taken up by affected brain cells and is enough to correct brain HS storage and• neuro inflammation to near normal levels and completely corrects the hyperactive• behaviour in mice with Sanfilippo
- 234. Mucopolysaccharidosis Type IIIA potential gene therapy • � Mucopolysaccharidosis Type IIIA (MPSIIIA) is a metabolic disorder in which the body is missing an enzyme that is required to break down long chains of sugars known as• glycosaminoglycans• • � The glycosaminoglycans collect in the body and cause damage, particularly in the brain if not broken• • � Fàtima Bosch and colleagues at Universitat Autònoma de Barcelona in Spain developed a form of gene therapy to replace the enzyme that is missing in MPSIIIA• • � They injected the replacement gene into the cerebrospinal fluid that surrounds the brain and spinal cord• • � This study demonstrates that gene therapy can be delivered to the brain through the cerebrospinal fluid and suggests that this approach could potentially be used as a• therapy for MPSIIIA•
- 235. IS GENE THERAPY TOTALLY SAFE ?? • Although gene therapy is a promising treatment • option for a number of diseases (including inherited • disorders, some types of cancer, and certain viral • infections), the technique remains risky and is still • under study to make sure that it will be safe and • effective. • Gene therapy is currently only being tested for the • treatment of diseases that have no other cures
- 236. Technical Difficulties in Gene Therapy • Gene delivery: Successful gene delivery is not easy or • predictable, even in single-gene disorders. • For example, although the genetic basis of cystic fibrosis is well • known, the presence of mucus in the lungs makes it physically • difficult to deliver genes to the target lung cells. • Delivery of genes for cancer therapy may also be complicated • by the disease being present at several sites. • Gene-therapy trials for X-linked severe combined • immunodeficiency (X-SCID), however, have been more • successful
- 237. Problems with Gene Therapy Short Lived• Hard to rapidly integrate therapeutic DNA into genome and rapidly• dividing nature of cells prevent gene therapy from long time• Would have to have multiple rounds of therapy• Immune Response• new things introduced leads to immune response• increased response when a repeat offender enters• the gene might be over• -expressed (toxicity) Viral Vectors• patient could have toxic, immune, inflammatory response• also may cause disease once inside•
- 238. Continued… Multigene• Disorders Heart disease, high blood pressure, Alzheimer• ’s, arthritis and diabetes are hard to treat because you need to introduce more than one• gene
- 245. Updates on current advances in gene therapy • At present, the three main gene therapy strategies for treatment of cancer are application to oncolytic viruses, suicide-gene therapy and gene- based immunotherapy.
- 247. AGGRESSIVE GENE THERAPY FOR CANCER • When dealing with cancer we need to destroy the cancer cells, or at least inhibit their growth and division. • Several strategies have been used and may be classified as follows:- (a) Genetic replacement (b) Direct attack (c) Suicide (d) Immune provocation
- 248. (A) Gene replacement Gene replacement therapy for cancer is use in correcting hereditary defect The cancer is analyzed to identify the mutant gene(s) that are responsible. Oncogene or tumour suppression gene is then inserted into the cancer cells. For example, p 53 gene has been delivered to p53 deficient cancer cells. The delivery method is usually via an adenovirus vector, but sometimes liposomes have been used.
- 250. (B) Direct attack In the direct plan of attack, a gene that helps kill cancer cells is used. For example, the TNF gene encodes tumour necrosis factor. This is produced by white blood cells known as tumor infiltrating lymphocytes. The cells normally infiltrate into tumour where they release TNF, which is fairly effective at eradicating small cancers. To attack a large cancer that is out of control, TNF production must be increased. First the TNF gene is cloned. Then white blood cells are removed from the patient and cultured. Multiple copies of the TNF gene -or an improved TNF gene with enhanced activity-are introduced into white cells. Then the white cells are injected back into patient. •
- 251. (C) SUICIDE GENE THERAPY Suicide strategy is a hybrid of anticancer drug therapy with gene transfer therapy. • Suicide gene therapy begins by delivering a therapeutic gene into the cancer cells. The genes encodes an enzyme that will convert a nontoxic product into a toxic compound. Because non-cancerous cells do not have the suicide enzyme, they are not affected. The non-toxic prodrug, ganciclovir, is converted to its monophosphate by thymidine kinase. Because only the cancer cells have thymidine kinase, all the noncancerous cells are unaffected. Normal cellular enzyme then convert monophosphate to GCV-TP. This act as a DNA chain terminator. DNA synthesis is inhibited and the cell is killed.
- 253. (D) IMMUNE PROVOCATION A more indirect approach relies on the body’s natural defences. Our immune systems are effective at killing cancers, provided they identify them while still small. To survive, a cancer has to somehow evade the body’s immune survillance. In this approach, gene therapy inserts a gene that attract the attention of the immune system to the tumor cells. A related approach is to use cytokines. These are short proteins that attract immune cells and stimulate their division and development. The genes for several cytokines of the interleukin family (especially IL2, IL4 and IL12) have been used to provoke immune attack on cancer cells. •
- 254. SUCCESS CASES OF GENE THERAPHY
- 256. GENE THERAPY REDUCES PARKINSON’S DISEASE SYMPTOMS • � It significantly improved the weakness of the • symptoms such as tremors, motor skill problems, and • rigidity. • � Main- overactive brain region: the subthalamic • nucleus should be introduced with gene. • � That would produce GABA—an inhibitory • chemical—then they could potentially quiet that • brain region and alleviate tremors.
- 257. HOW IT WORKS ?? • Done with local anesthesia, used a harmless, • inactive virus [AAV-2 GAD]. • Deliver the GAD gene into patient’s • subthalamic nucleus. • The gene instructs cells to begin making • GABA neurotransmitters to re-establish the • normal chemical balance that becomes • dysfunctional as the disease progresses.
- 259. REFERENCES • Jogdand, S.N. (2006). Gene Biotechnology. Himalaya Publishing House. Mumbai, India. 2nd ed., p 237-249. • Chen, C.A. & Okayama, H. (1988). Calcium phosphate-mediated gene transfer: a highly efficient trasfection system for stably transforming cells with plasmid DNA. Biotechniques., • Watwe, R.M. & Bellare, J.R. (1995). Manufacture of liposomes a review. Curr. Sci. India., . Nicolau, C., Legrand, A.& Grosse, E. (1987). Liposomes as carriers for in vivo gene transfer and expression. Method Enzymol., 149: • 157-176. • lies, M.A. & Balaban, A.T. (2001). Recent developments in cationic lipid-mediated gene delivery and gene therapy. • Expert Opin. Ther. Patents., • Reece, R.J. (2004). Analysis of Genes and Genomes. John Wiley and Sons Ltd. • Johnston, S.A. & Tang, D.C. (1994). Gene gun transfection of animal cells and genetic immunization. Methods Cell • Biol., 43 •Klein, T.M., Arentzen, R., Lewis, P.A. & Fitzpatrickmcelligott, S. (1992). Transformation of microbes, plants and animals by particle bombardment. Biotechnology • King, R. (2004). Gene delivery to mammalian cells by microinjection. Methods Mol. Biol., • David B. Burr; Matthew R. Allen (11 June 2013). Basic and Applied Bone Biology. Academic. p. 157. ISBN 978-0-12- 391459-0. • Retrieved 15 July 2013. • ^ Juan Carlos Lacal; Rosario Perona; James Feramisco (11 June 1999). Microinjection. Springer. p. 9. ISBN 978-3-7643- 6019-1. Retrieved 13 July 2013.
